How to run a development environment for the backend
This page describes a list of steps to follow to run a development environment for the backend locally
We assume in this guide:
- That you have access to a shell Terminal, either because you are running on a Linux/Mac operating system or Windows wth WSL 2 installed
You will use this same backend environment for ontologies_api, ontologies_linked_data, and goo projects
Steps to follow
Run the backend infrastructure using VirtualBox
We will use a virtual machine with a nat port forwarding of all ports needed to run the ontologies_api project locally. Port forwarding will make them all accessible from the localhost
Install the appliance on VirtualBox
First get the appliance and run it on VirtualBox
Configure port forwarding
Then on VirtualBox go to Settings > Network, and change the adapter to NAT and click on Advanced then Port Forwarding. Where you add the following ports to forward:
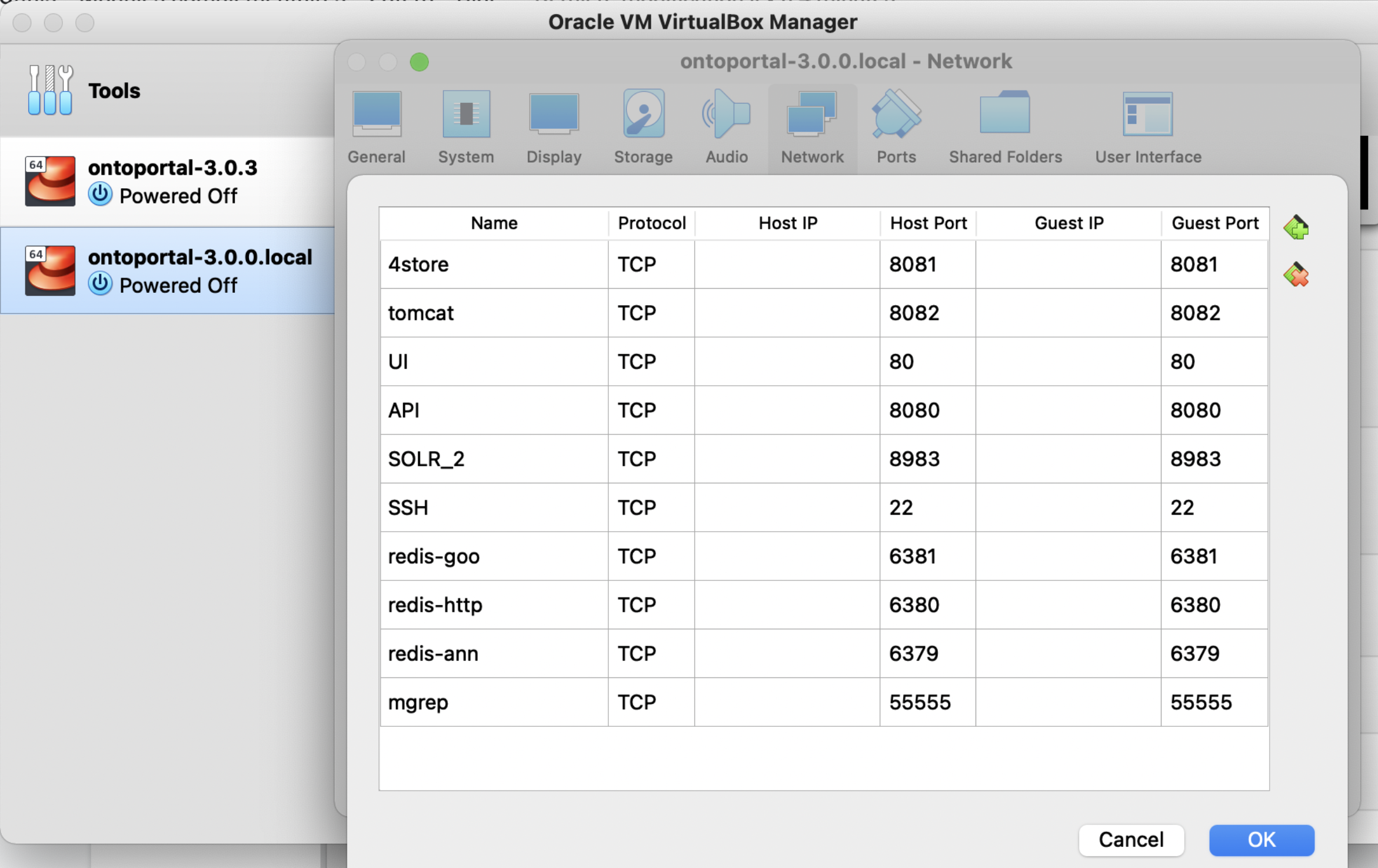
If mapping SSH to port 22 didn’t work, you can use a different port port (e.g 2222)
Finally start the appliance and connect to the appliance with this command line: ssh centos@127.0.0.1
Important: if you can’t establish an ssh connection with the appliance ,make sure to run first the SSH service in the appliance, using this command: sudo systemctl enable sshd
Important: to make the port forwarding of redis and 4store work, be sure to disable first the firewall of the appliance, after connecting to the appliance via ssh using the command: sudo systemctl stop iptables
Run the code locally
Fork and clone backend source codes
First fork the ontologies_api repository. More details about how to can be found in the Github official doc
Then clone your forked repository locally
git clone https://github.com/your_organization/ontologies_api.git
The master branch represent the Bioportal version, if you want to have the Agroportal version. You have to merge the ontoportal-lirmm branch to your master, by doing so git merge ontoportal-lirmm master
Create the config file config/environments/development.rb
A sample file of config/environments/development.rb can be found here: https://github.com/ontoportal/ontologies_api/blob/master/config/environments/config.rb.sample
In command line do
cp config/environments/config.rb.sample config/environments/development.rb
Copy the content bellow in the config/environments/development.rb
$HOSTNAME = $UI_HOSTNAME = "ontoportal.local"
$REST_HOSTNAME = "localhost"
$REST_PORT = 9393
$REST_URL_PREFIX = "http://#{[$REST_HOSTNAME, $REST_PORT].compact.join(':')}/"
$DATADIR ||= '/srv/ontoportal/data'
GOO_BACKEND_NAME = '4store'
GOO_PORT = GOO_BACKEND_NAME.include?('AG') ? 10035 : 8081
GOO_PATH_QUERY = GOO_BACKEND_NAME.include?('AG') ? '/repositories/ontoportal' : '/sparql/'
GOO_PATH_DATA = GOO_BACKEND_NAME.include?('AG') ? '/repositories/ontoportal/statements' : '/data/'
GOO_PATH_UPDATE = GOO_BACKEND_NAME.include?('AG') ? '/repositories/ontoportal/statements' : '/update/'
begin
# For prefLabel extract main_lang first, or anything if no main found.
# For other properties only properties with a lang that is included in main_lang are used
Goo.main_lang = ["en", "eng", "fr", "fre"]
Goo.use_cache = false
rescue NoMethodError
puts "(CNFG) >> Goo.main_lang not available"
end
begin
LinkedData.config do |config|
config.goo_host = 'localhost'
config.goo_port = "#{GOO_PORT}"
config.goo_backend_name = "#{GOO_BACKEND_NAME}"
config.goo_path_query = "#{GOO_PATH_QUERY}"
config.goo_path_data = "#{GOO_PATH_DATA}"
config.goo_path_update = "#{GOO_PATH_UPDATE}"
config.rest_url_prefix = "#{$REST_URL_PREFIX}"
config.ui_host = "#{$UI_HOSTNAME}"
config.search_server_url = 'http://localhost:8983/solr/term_search_core1'
config.property_search_server_url = 'http://localhost:8983/solr/prop_search_core1'
config.repository_folder = "#{$DATADIR}/repository"
config.replace_url_prefix = true
config.enable_security = true
config.enable_slices = true
# Caches
Goo.use_cache = false
config.goo_redis_host = "localhost"
config.goo_redis_port = 6381
config.enable_http_cache = true
config.http_redis_host = "localhost"
config.http_redis_port = 6380
# PURL server config parameters
config.enable_purl = false
config.purl_host = "purl.example.org"
config.purl_port = 80
config.purl_username = "admin"
config.purl_password = "password"
config.purl_maintainers = "admin"
config.purl_target_url_prefix = "http://example.org"
# Email notifications
config.enable_notifications = true
config.email_sender = "notifications@test.com" # Default sender for emails
config.email_override = "notifications@test.com" # all email gets sent here. Disable with email_override_disable.
config.email_disable_override = true
config.smtp_host = "smtp.lirmm.fr"
config.smtp_port = 25
config.smtp_auth_type = :none # :none, :plain, :login, :cram_md5
config.smtp_domain = "lirmm.fr"
# Emails of the instance administrators to get mail notifications when new user or new ontology
config.admin_emails = []
# Ontology Google Analytics Redis
# disabled
config.ontology_analytics_redis_host = "localhost"
config.ontology_analytics_redis_port = 6379
# Used to define other bioportal that can be mapped to
# Example to map to ncbo bioportal : {"ncbo" => {"api" => "http://data.bioontology.org", "ui" => "http://bioportal.bioontology.org", "apikey" => ""}
# Then create the mapping using the following class in JSON : "http://purl.bioontology.org/ontology/MESH/C585345": "ncbo:MESH"
# Where "ncbo" is the namespace used as key in the interportal_hash
config.interportal_hash = {
"agroportal" => {
"api" => "http://data.agroportal.lirmm.fr",
"ui" => "http://agroportal.lirmm.fr",
"apikey" => "1cfae05f-9e67-486f-820b-b393dec5764b"
},
"ncbo" => {
"api" => "http://data.bioontology.org",
"ui" => "http://bioportal.bioontology.org",
"apikey" => "4a5011ea-75fa-4be6-8e89-f45c8c84844e"
},
"sifr" => {
"api" => "http://data.bioportal.lirmm.fr",
"ui" => "http://bioportal.lirmm.fr",
"apikey" => "1cfae05f-9e67-486f-820b-b393dec5764b"
}
}
end
rescue NameError
# puts '(CNFG) >> LinkedData not available, cannot load config'
end
begin
Annotator.config do |config|
config.mgrep_dictionary_file = "#{$DATADIR}/mgrep/dictionary/dictionary.txt"
config.stop_words_default_file = './config/default_stop_words.txt'
config.mgrep_host = 'localhost'
config.mgrep_port = 55555
config.mgrep_alt_host = 'localhost'
# secondary mgrep instance is not configured for appliance. routing all requestes to the primary mgrep
config.mgrep_alt_port = 55555
config.annotator_redis_host = 'localhost'
config.annotator_redis_port = 6379
end
rescue NameError
# puts '(CNFG) >> Annotator not available, cannot load config'
end
begin
OntologyRecommender.config do |config|
end
rescue NameError
# puts '(CNFG) >> OntologyRecommender not available, cannot load config'
end
begin
LinkedData::OntologiesAPI.config do |config|
config.enable_unicorn_workerkiller = true
config.enable_throttling = false
config.http_redis_host = LinkedData.settings.http_redis_host
config.http_redis_port = LinkedData.settings.http_redis_port
config.restrict_download = []
#config.ontology_rank = ""
end
rescue NameError
# puts '(CNFG) >> OntologiesAPI not available, cannot load config'
end
begin
NcboCron.config do |config|
config.redis_host = Annotator.settings.annotator_redis_host
config.redis_port = Annotator.settings.annotator_redis_port
config.enable_ontology_analytics = false
config.search_index_all_url = 'http://localhost:8983/solr/term_search_core2'
config.property_search_server_index_all_url = 'http://localhost:8983/solr/prop_search_core2'
config.ontology_report_path = "#{$DATADIR}/reports/ontologies_report.json"
config.enable_spam_deletion = false
end
rescue NameError
puts '(CNFG) >> NcboCron not available, cannot load config'
end
Install dependencies
You will need to install Ruby, for the version see the RUBY_VERSION argument
Link your local ontologies_linked_data and goo code to ontologies_api project
If you want to use your local code for ontologies_linked_data and goo you will need also to fork and clone them from the Ontoportal repositories
In addition, you will need to configure the bundler to use them, by doing like below
cd put_here_the_path_to_your_local_ontologies_api_code
bundle config local.goo put_here_the_path_to_your_local_goo_code
bundle config local.ontologies_linked_data put_here_the_path_to_your_local_ontologies_linked_data_code
Run the ontologies_api server
bundle exec shotgun --env=development